Current Research Focus
Epigenetic effects of copper oxide nanoparticles leading to teratogenicity in zebrafish
Nano-copper oxides are a versatile inorganic material. As a result of
their versatility, the immense applications and usage end up in the
environment causing a concern for the lifespan of various beings. The
ambiguities surround globally on the toxic effects of copper oxide
nanoparticles (CuO-NPs). Hence, we endeavored to study the sub-lethal
acute exposure effects on the developing zebrafish embryos. The 48 hpf
LC50 value was about 64 ppm. The sub-lethal dose of 40 and 60 ppm
led to the developmental anomalies in addition to oxidative stress
in the developing embryos.
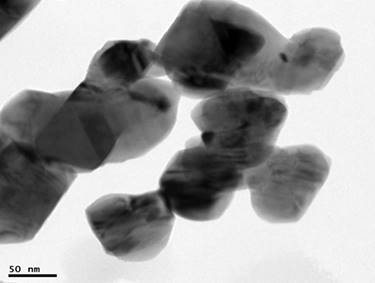
TEM image of CuO-NPs
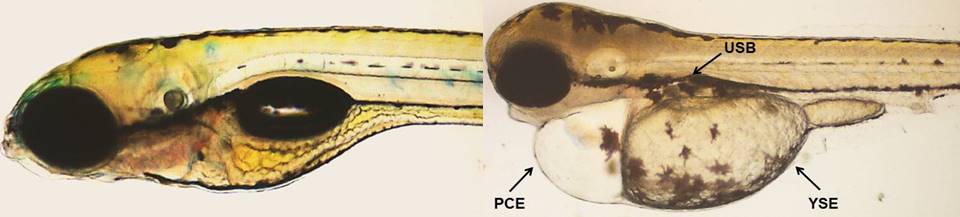
Normal (Control) and malformed zebrafish larvae exposed to CuO-NPs. USB - Uninflated swim bladder,
PCE - Pericardial edema, YSE - Yolk sac edema
With this valuable data, we are now
exploring the epigenetic mechanisms involved in the induction of
teratogenicity caused by CuO-NPs. The objectives are:
▢ Investigation of epigenetic modifications of DNA and changes in the
transcriptome in CuO-NPs exposed zebrafish.
▢ Determination of microRNAs in teratogenicity induction in CuO-NPs
exposed zebrafish.
▢ Elucidation of histone modifications (methylation, acetylation, and
phoshorylation) in zebrafish on treatment with CuO-NPs.
Role of fisetin on the interrelationship between autophagy and apoptosis in hepatocellular carcinoma
Hepatocellular carcinoma (HCC) is the one of the leading liver cancers
in many countries and is the fifth most common cancer worldwide. Though
HCC management has improved in recent years through the advances in
prevention and treatment, it still requires the effective mechanism
based approaches. When cancer cells are treated with different drugs,
the interactions between autophagy and apoptosis will have profound
effects on the outcome of the tumor cell. The cell may undergo autophagy
by blocking apoptosis or by apoptosis when autophagy is inhibited. The
discovery of cross link between the apoptosis machinery and autophagic
machinery lead to a situation in better management of HCC.
Fisetin (3,7,3',4'- Tetrahydroxyflavone) is a plant secondary metabolite
found in many fruits and vegetables such as tomatoes, onions, grapes,
strawberry, persimmon, cucumber and apples which are regularly consumed
in the human diet. This natural flavonoid reportedly exhibits antioxidant,
neurotrophic, anti-inflammatory and anti-cancer effects. These properties
necessitate fisetin to be further studied for its potential as a
chemopreventive agent in HCC. Fisetin may afford chemopreventive as well
as cancer chemotherapeutic effects against HCC. However, the effects of
fisetin on HepG2 cells have not yet been investigated.
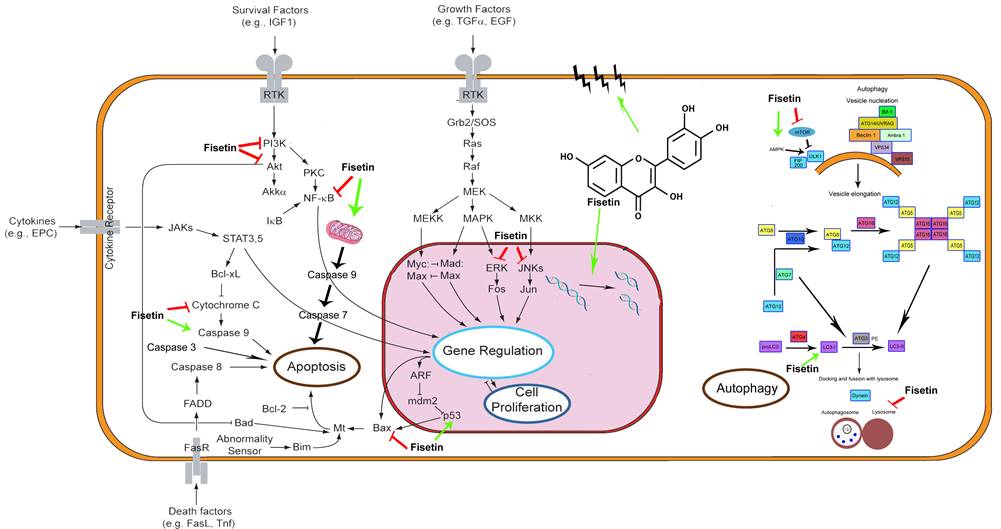
Proposed mechanism of fisetin on interrelationship
between autophagy and apoptosis in hepatocellular carcinoma
In our lab, we
investigate the effect of fisetin on apoptosis and autophagy in HepG2
cells with following objectives:
▢ Determination of the specific mode of action of fisetin on multiple
signaling pathways leading to cell death using HepG2 cells.
▢ Establish the role of fisetin in apoptosis and autophagy signaling
pathways.
▢ Exploration of the crosslink between apoptosis and autophagy on
fisetin treated HepG2 cells.
Identifcation, validation, and analysis of molecular activators and antioxidant response
elements in the Keap1-Nrf2-ARE pathway in zebrafish
The Keap1-Nrf2-ARE system serve as a permier defense mechanism to
curb oxidative stress, thereby safeguard cells and organs. Nrf2 -
the principal master regulator of the cellular defense system. The
Keap1-Nrf2-ARE pathway is regulated by the Keap1-Nrf2 protein-protein
interaction (PPI). Keap1 is a negative regulator of this pathway and
in zebrafish, there are two types of Keap1: Keap1a and Keap1b.
Recently, we identified top compounds that disrupts both Keap1a and
Keap1b interaction with Nrf2. These potential can serve as safer Nrf2
activators due to their non-covalent disruption of Keap1-Nrf2 PPI.
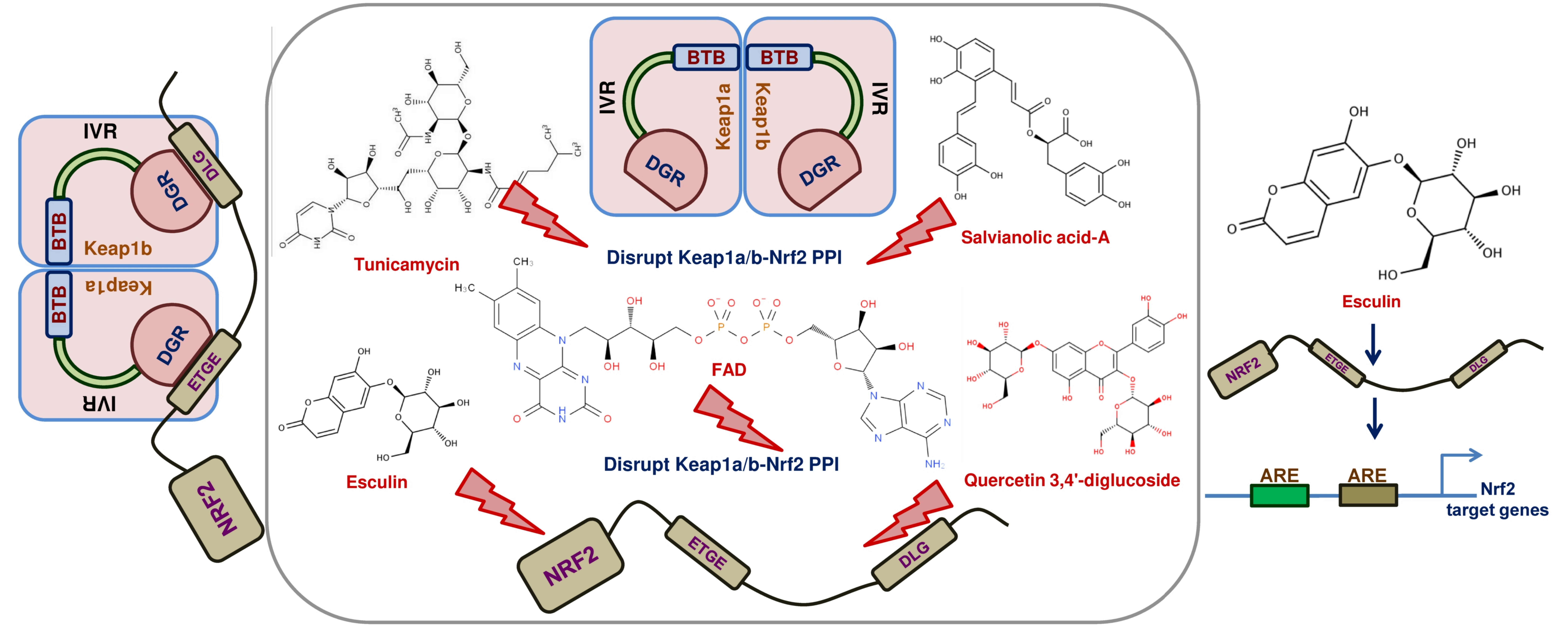
Molecular activators that disrupt the interaction
between Keap1a/b and Nrf2 in zebrafish
Nrf2 - a transcription factor which recognise specific DNA sequences
upstream of the genes. These regulatory elements are called antioxidant
response elements (ARE), which define outcome of gene expression.
AREs are only a fraction of the total genome in zebrafish. Identifying
these AREs is a daunting task. In the post-genomic era, deciphering the
Nrf2 binding sites - AREs is an essential task that underlies and
governs this Keap1-Nrf2-ARE pathway. Recently, we identified AREs in
the genome of zebrafish through a pattern search algorithm and analyzed
using computational tools available online. AREs are identified within
30 kb upstream from the transcription start site of antioxidant genes,
mitochondrial genes, and all the known protein coding genes in the
zebrafish genome. The findings revealed that TGAG/CNNNTC ARE motifs were
found to be abundant than TCAG/CNNNGC ARE motifs in the antioxidant genes
and in all protein coding genes of zebrafish. Our results help to
understand the dynamic complexity of the Nrf2-ARE system in zebrafish.
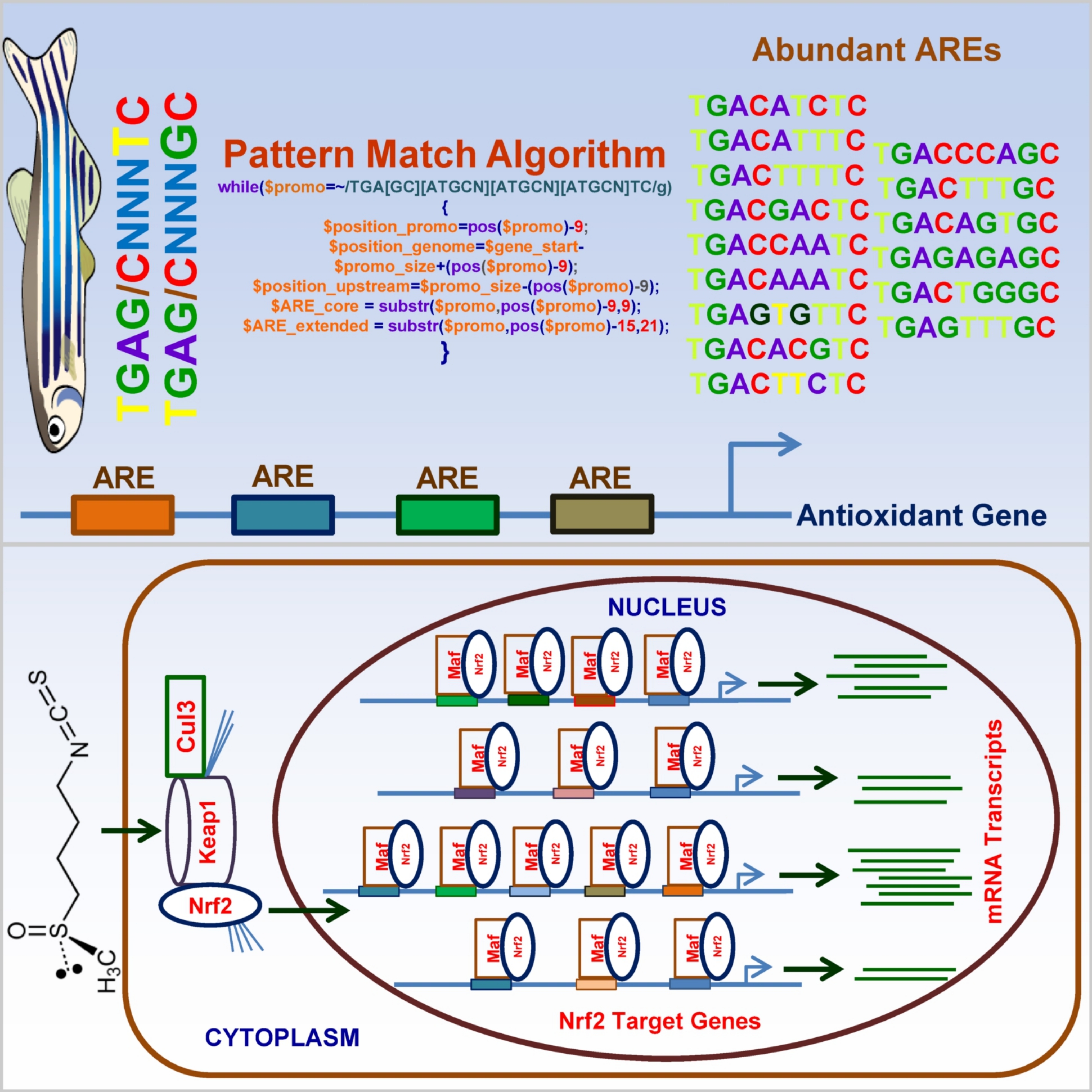
Abundant AREs identified in zebrafish and their
roles in the transcription of Nrf2-target genes
In order to disseminate and share the identified AREs in zebrafish. We
have developed a database named, Zebrafish Antioxidant Response Element
Database (ZFARED) www.zfared.buc.edu.in - an online open-access resource for
AREs in zebrafish.

ZFARED: Zebrafish antioxidant response element
database
Based on these discoveries, our lab is interested in understanding
how Keap1-Nrf2-ARE signaling pathway regulates gene expression through
following objectives:
▢ Identification and validation of compounds that disrupt the Keap1-Nrf2 PPI in vivo
in zebrafish embryo/larva model.
▢ Functional characterization of identified AREs using molecular
genetic approaches in zebrafish.
Research Grants
Ongoing Projects
▢ A grant of INR.46,19,000/- towards the project entitled, "Identification, validation,
analysis and characterization of molecular activators and antioxidant response elements
of Keap1/Nrf2/ARE signaling pathway in zebrafish" from March 2019 to March 2022 by
Empowerment and Equity Opportunities for Excellence in
Science, Department of Science and Technology – Science and Engineering Research
Board, New Delhi.
Completed Projects
▢ A grant of INR.11,00,000/- toward the project entitled, “Role of fisetin on the
interrelationship between autophagy and apoptosis in hepatocellular carcinoma” from
March 2016 to March 2018 by Empowerment and Equity Opportunities for Excellence in
Science, Department of Science and Technology – Science and Engineering Research
Board, New Delhi.
▢ A grant of INR.20,20,000/- towards the project entitled, “Catecholaminergic activity
enhancer compounds for performance enhancement in vitro and in vivo models” from April
2014 to March 2016 by Defence Research and Development Organization, New Delhi.
▢ A grant of INR.33,77,625/- towards the project entitled, “Elucidation the molecular
of necroptosis in fluoride induced neurotoxicity: Therapeutic potential of Naringenin
through Nf?B and Nrf2 signalling pathways” from December 2015 to December 2018 by
Department of Science and Technology – Science and Engineering Research Board, New Delhi.
▢ A grant of INR.13,22,380/- towards the project entitled, “Effects of tamarind
seed coat extract on fluoride induced apoptosis in MRC-9 cells” from February
2010 to January 2013 funded by University Grants Commission, New Delhi.
|


